Sponsored by HORIBAJun 14 2018
Surface Plasmon Resonance imaging (SPRi) is a technique which can help monitor and study interactions between biomolecules without the use of a label, using optical real-time detection. It uses imaging capability to allow the whole working area to be seen and to permit multiplex analysis.
What Does Multiplex Mean?
A multiplex analysis is one in which a variety of ligands can be immobilized using one SPRi-Biochip, to be analyzed for multiple parameters such as their concentration, immobilization, and pH. This adds convenience to the comparison, ranking and selection of molecules for further study and development.
What is the Principle of SPRi?
SPRi technology is aimed at measuring the changes in the refractive index measured at the surface of the SPRi-Biochip since such changes are a function of the mass variations. This can be used for the real-time identification of molecular interactions, their affinity, and the concentration of various analytes involved.
What Kind of Data is Collected?
SPRi technology yields a rather comprehensive picture of the interaction between biomolecules, including the degree of specificity, kinetics of the reaction and the affinity, which is useful to characterize a sample solution with respect to the quantity of molecules.
What Interactions can be Analyzed?
The samples that can be analyzed in this way comprise of a huge library of carbohydrates, proteins and peptides, nucleic acids, cells, bacteria, polymers, and small organic molecules, among others.
How can SPRi be Used to Study Molecular Interactions?
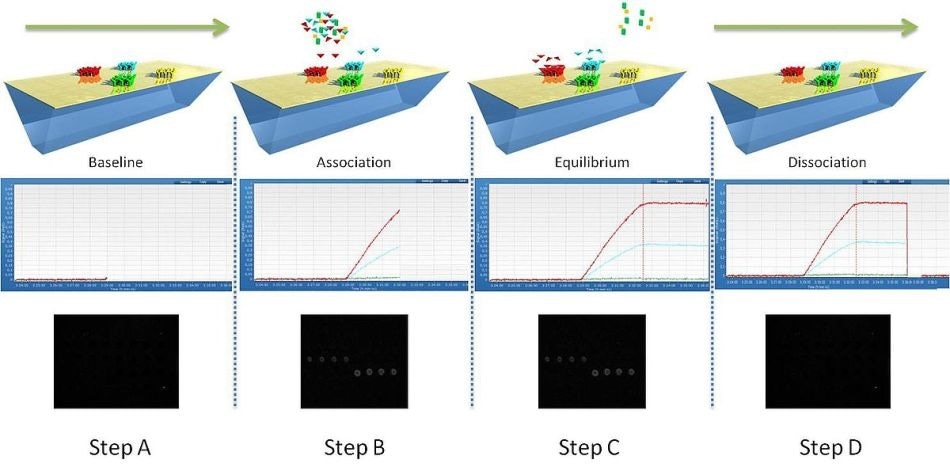
Figure 1. Monitoring of molecular interactions by SPRi.
The usefulness of SPRi lies in its ability to detect the changes that occur in the refractive index at the SPRi-Biochip surface. This is related to any event which leads to a mass accumulation or binding of biomolecular domains. The change in the refractive index results in a corresponding shift in the resonance angle position. The changes in reflectivity that occur with a fixed angle (called the working angle) over time are monitored by SPRi as illustrated in Figure 1.
In the first step, ligand immobilization occurs on the surface of the functionalized SPRi-Biochip in an array.
In the second step, the dissolved sample is injected into the flow cell. This initiates the conditions for molecular binding which in turn shifts the plasmon curves, causing an enhancement in the reflectivity. The kinetic curves, also called sensor grams, clearly depict how reflectivity varies with time in the association phase. An SPRi difference image can also be generated to monitor this interaction, with the white spots being associated with the areas of the SPRi-Biochip where interactions are occurring.
In the next step, the solution containing the sample is discharged from the flow cell. This leads to dissociation of the complex formed by the ligand and analyte, resulting in a shift of the plasmon curve and a reduction in the surface reflectivity. The sensor grams show the dissociation phase by the changes in reflectivity plotted against time. The SPRi difference image shows this as darkening of the interaction areas shown as spots on the image.
In the final step, complete dissociation of all the ligand-analyte complexes has occurred. This may be facilitated if required, using a regeneration solution. This is shown by the return of the plasmon curves and the kinetic curves to the state they were in at the beginning, while the SPRi difference image becomes black again.

This information has been sourced, reviewed and adapted from materials provided by HORIBA.
For more information on this source, please visit HORIBA.