Protein characterization can be an extensive field of study covering a broad range of analytical methods and techniques. Although biochemists have made significant advances in protein analysis, the process of identifying and purifying novel proteins remains a formidable task. This is largely due to the lack of a universal solution for macromolecules with divergent aggregation states, sizes, charges, and structures.
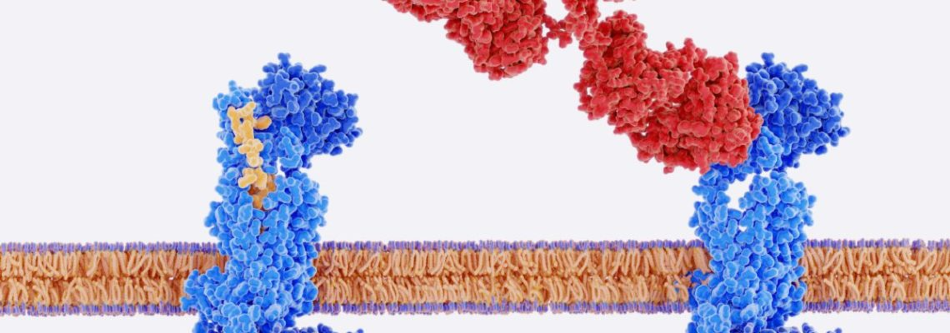
Image Credit: Jordi Labs
The sheer magnitude and complexity of protein characterization stems from the almost endless amount of pathways for protein expression. All proteins are comprised of chains of the same 21 amino acid residues of variable concentrations that assemble in near-infinite arrangements. They can then be folded into three-dimensional (3D) structures that also differ in size. The protein characterization process is further complicated by the fact that they are never present in isolation: a single cell may contain as many as 10,000 different proteins.
How do biochemists even begin to perform good, accurate protein analysis and characterization?
Purifying Proteins
To properly characterize a single protein, chemists must initially isolate it from a sample via a purification method and identify it by any number of defining characteristics. As previously stated, to perform protein characterization biochemists utilize a variety of analytical methods ranging from the routine to the experimental.
The acquisition of proteins for analysis commences when a sample is selected and fractionated. For example, cells can be lysed and the required sample material can be extracted through differential centrifugation. An increasingly pure supernatant is separated from larger sample debris. Even after several passes in a centrifuge, it is probable that the supernatant will contain thousands of distinct proteins.
Purification of the protein of interest takes place by subjecting the sample to a separation technique based on intrinsic chemical, electrical, or physical properties. Chromatography (affinity, gel-filtration, ion-exchange, HPLC, etc.) allows for highly-selective separations of sample material based on various characteristic properties, making it one of the most common technologies used in protein purification. Jordi Labs assists companies in understanding the structure of the protein molecule.
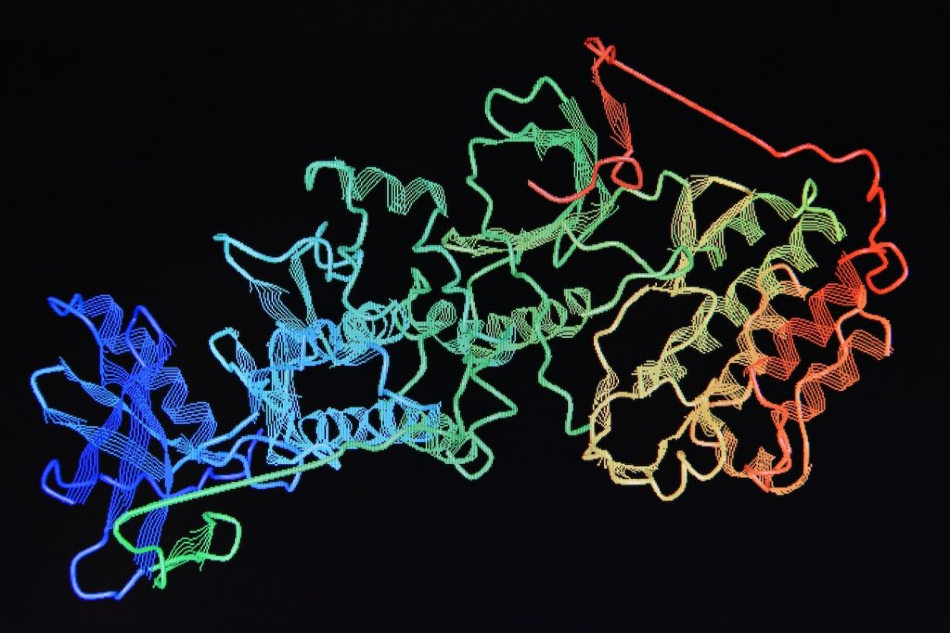
Image Credit: Jordi Labs
Identifying Proteins
Once acquisition of a highly pure protein has been finalized, biochemists can then commence the characterization of molecular weight, structure, composition, and purity/impurity. These objectives are often interconnected and identification of specific values is best achieved when performed in a particular order. For example, the molecular weight of a protein chain is anticipated based on the amino acid composition, which is determined by chemical methods.
To determine the precise amino acid sequence of proteins chemically, its peptide backbone is cleaved via a number of potential methods such as tryptic digestion, where residues are cleaved sequentially from the protein chain and determined via high-pressure liquid chromatography (HPLC) and mass spectrometry. This is the most common method for chemically identifying an individual protein’s sequence.
This is just one clear example of how proteins are identified and what is discovered relative to their chemical structure. Other examples include:
- Aggregation state
- Extinction coefficient (molar absorptivity)
- Homo- and heterogeneity
- Molecular weight
- Protein Modifications (glycosylation, PEGylation, etc.)
- Purity/impurity
- Spectroscopic characteristics
Jordi Labs provides supreme protein characterization and analysis utilizing a suite of well-established tools. If you have novel protein analysis needs and think you might benefit from Jordi’s support, simply contact a member of the team today.

This information has been sourced, reviewed and adapted from materials provided by Jordi Labs.
For more information on this source, please visit Jordi Labs.