Dendrimers are highly branched macromolecules with a tree-like architecture that grows outward from a central core.
Lower-generation dendrimers feature simpler structures that are useful in routine applications, while higher generations have more terminal groups and improved reactivity, making them highly useful for advanced applications such as molecular recognition, drug delivery, and nanotechnology.
Dendrimers exhibit high molecular density and a small hydrodynamic volume due to their compact structure. This often causes them to elute later in Size Exclusion Chromatography (SEC).
Traditional approaches to calibration are based on elution time, but these typically underestimate molecular weight. Absolute molecular weight can be determined from scattering intensity rather than retention time by integrating light scattering detection within the BeSEC. This approach ensures accurate characterization of dendritic materials.
Experimental
The study presented here used an SEC system equipped with light-scattering (LS) and refractive index (RI) detectors. The BeSEC LS2 from Bettersize Instruments features a light-scattering detector with 90 ° and 7 ° angles.
The BeSEC workstation combines light scattering with RI or UV signals to calculate molecular weight averages (Mn, Mw, and Mz) and distributions.
System Configuration
The system configuration employed was as follows:
- Detectors: Light Scattering (LS) + Refractive Index (RI)
- Column: Shodex GPC KF-806M
- Mobile phase: 0.02 M LiBr in DMF
- Flow rate: 0.7 mL per minute
- Injection volume: 100 μL
- Column temperature: 40 °C
- dn/dc: Calculated from sample concentration
Sample Preparation
Two dendrimer samples were analyzed: G5 and G7.5, corresponding to Generations 5 and 7.5, respectively.
Each sample was accurately weighed before being dispersed in DMF (10–20 mg/mL). It was then stirred until clear and filtered through a 0.22 μm PTFE syringe filter. Finally, the sample was transferred into vials to enable autosampler injection.
Results and Discussion
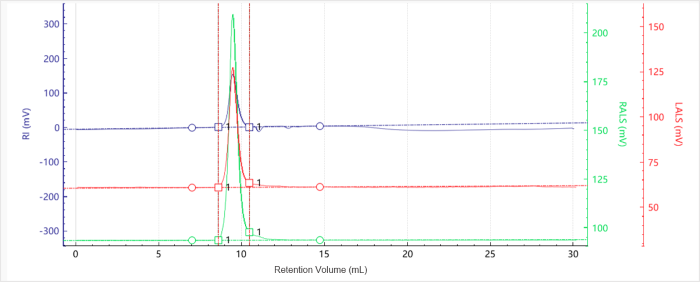
Figure 1. Elution profiles of the multi-detector signals for Sample G5. Image Credit: Bettersize Instruments.
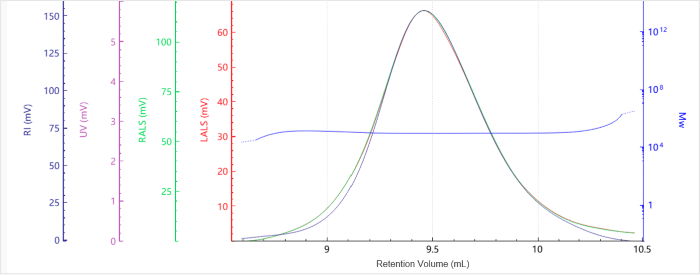
Figure 2. Elution profile of the molecular weight for Sample G5. Image Credit: Bettersize Instruments.
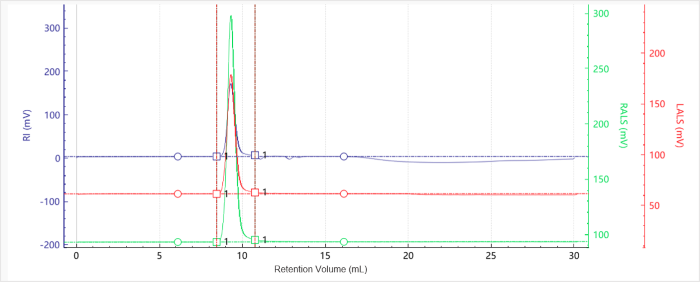
Figure 3. Elution profiles of the multi-detector signals for Sample G7.5. Image Credit: Bettersize Instruments.
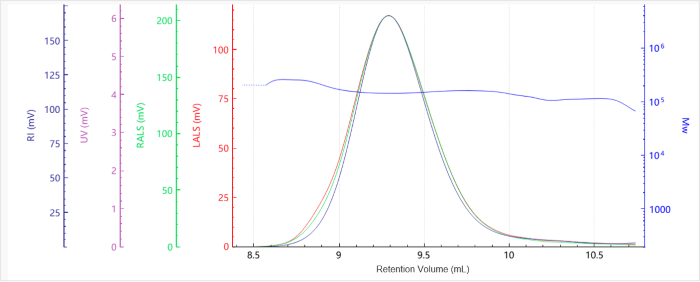
Figure 4. Elution profile of the molecular weight for Sample G7.5. Image Credit: Bettersize Instruments.
Figures 1 to 4 show multi-detector chromatograms:
- RI signals are shown in blue
- Right-angle light scattering (RALS) signals are shown in green
- Low-angle light scattering (LALS) signals are shown in red
The baselines were flat with excellent signal quality and minimal noise. It was noted that molecular weight profiles varied only slightly with elution volume, highlighting the presence of narrow molecular weight distributions.
Both dendrimer samples eluted early. This occurred close to the solvent peak at approximately 11 minutes, highlighting their particularly small hydrodynamic volume.
Using conventional calibration based solely on RI would result in species in this region being assigned molecular weights of only a few thousand Daltons, significantly below their actual values.
Table 1. Molecular weight results of dendrimer samples. Source: Bettersize Instruments.
| No. |
Mn (Da) |
Mw (Da) |
Mz (Da) |
Mw/Mn |
| Sample G5 |
102,233 |
103,542 |
106,627 |
1.0128 |
| Sample G7.5 |
156,451 |
158,750 |
162,323 |
1.0149 |
Table 1 confirms that the weight-average molecular weight increases with dendrimer generation, with both samples exceeding 100 kDa. Light-scattering detection delivers reliable absolute molecular weight measurements by accurately capturing this trend.
Conclusion
The study presented here used the BeSEC LS2 to confirm that dendrimer molecular weights consistently rise with generation.
The compact nature of dendritic structures is illustrated by their narrow distributions and early elution. Unlike traditional calibration methods, light scattering provides accurate molecular weight characterization, avoiding significant underestimation and facilitating precise analysis of dendritic materials.
Acknowledgments
Produced from materials originally authored by Zhibin Guo from Bettersize Instruments.

This information has been sourced, reviewed and adapted from materials provided by Bettersize Instruments.
For more information on this source, please visit Bettersize Instruments .